◀ GH2_1
◀ GH53
◀ unk
◀ SusD
◀ SusC
◀ unk
◀ HTCS
Martens EC, Chiang HC, Gordon JI
Cell host & microbe, 4(5):447-57 (2008)
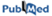
Recognition and degradation of plant cell wall polysaccharides by two human gut symbionts.
Martens EC, Lowe EC, Chiang H, Pudlo NA, Wu M, McNulty NP, Abbott DW, Henrissat B, Gilbert HJ, Bolam DN, Gordon JI
PLoS biology, 9(12):e1001221 (2011)
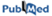
Dietary pectic glycans are degraded by coordinated enzyme pathways in human colonic Bacteroides.
Luis AS, Briggs J, Zhang X, Farnell B, Ndeh D, Labourel A, Baslé A, Cartmell A, Terrapon N, Stott K, Lowe EC, McLean R, Shearer K, Schückel J, Venditto I, Ralet MC, Henrissat B, Martens EC, Mosimann SC, Abbott DW, Gilbert HJ
Nature microbiology, 3(2):210-219 (2017)
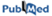
| Protein | Modularity | CAZy function | Genome Sequencing description | Pfam domains | ||
| BT4667 | ◀ GH2_1 [EC] |
EC3.2.1.23 | b-galactosidase |  |
beta-galactosidase | PF02837 PF00703 PF02836 PF16355 PF18565 |
| BT4668 | ◀ GH53 [EC] |
EC3.2.1.89 | endo-b-1,4-galactanase |  |
arabinogalactan endo-1,4-beta-galactosidase | PF07745 |
| BT4669 | ◀ unk |
hypothetical protein | PF14292 PF17138 PF17141 | |||
| BT4670 | ◀ SusD |
putative outer membrane protein, probably involved in nutrient binding | PF07980 | |||
| BT4671 | ◀ SusC |
putative outer membrane protein, probably involved in nutrient binding | PF13715 PF07715 PF00593 | |||
| BT4672 | ◀ unk |
hypothetical protein | ||||
| BT4673 | ◀ HTCS |
two-component system sensor histidine kinase | PF07494 PF07495 PF00512 PF02518 PF12833 | |||
| Predicted PUL 84 | ◀ GH2_1 ◀ GH53 ◀ unk ◀ SusD ◀ SusC ◀ unk ◀ HTCS unk ▶ PL13 ▶ |